B-Epitope Prediction Tools

EpiQuest-B is a program predicting and ranging domains of linear protein sequences according to their capacity to elicit humoral immune response. The program allows in silico prediction of immunodominant epitopes by assessing their potential antigenicity. This means that it does it equally for linear peptides irrespective of their exposure at the surface of the mature protein molecule and without consideration of post-processing modifications, such as glycosylation, phophorylation, etc.
The algorithm used in EpiQuest-B is based on the assumption that antigenicity of an epitope is defined by the nearest neighbors of amino acids in the sequence. The algorithm is based on the data on the contest for >40.000 individual amino acids in epitope sequences (strong, weak and negative) and tested the usability using >2000 individual epitopes with known immunization data from the immunizations performed in Aptum Biologics or its predecessor companies.

The algorithm of EpiQuest B calculates the probability of an aminoacid to be a part of an antigenic peptide sequence on the basis of aminoacids comprising its context.
Convenient Layout and Output
New layout of EpiQuest-B (EpiQuest v4.1 and later) allows to perform the antigenicity profiling of the protein sequences along with other types of analysis (accessibility, complexity, and the other ones). It also allows you to control the sensitivity of the analysis, focusing the output on only highly antigenic epitopes by adjusting the settings of the analysis.
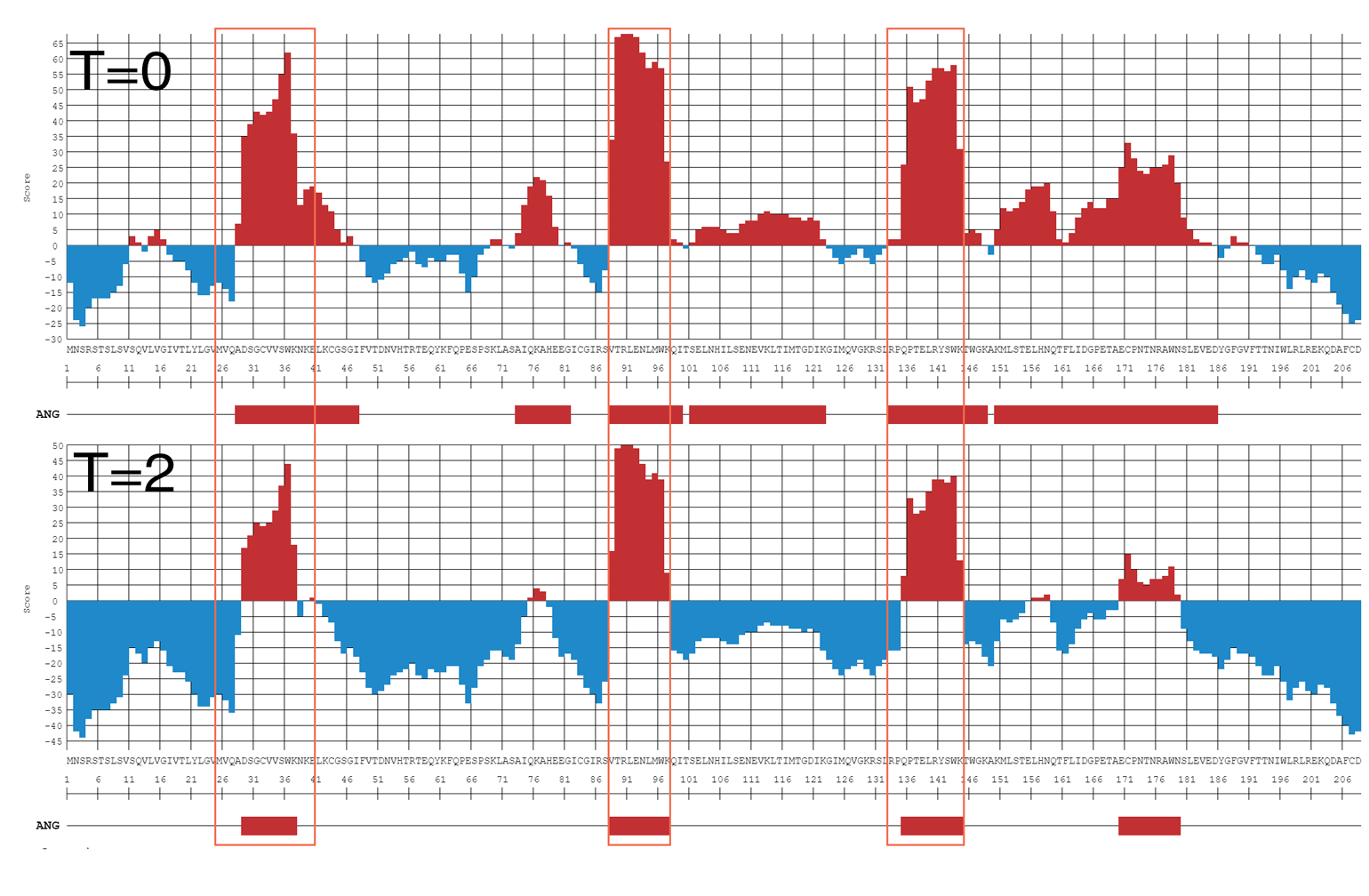
In this example adjustment of the Threshold (Th) allows to limit the detected epitopes to the immunodominant ones (the location of the actual immunodominant epitopes is indicated by the red frames). See more examples here.
Selecting optimal epitopes
For making a strong specific antibody recognizing with high affinity the target protein the selected epitope should be 1) highly antigenic, 2) exposed at the surface of the mature foolded target protein, 3) be unique, that is present only in this protein. There are EpiQuest programs developed exactly to find sigh epitope.
EpiQuest-B is program that detects and correctly predicts immunodominant epitopes, that is linear sequences capable of eliciting strong humoral response in immunized animal.
EpiQuest-A reviews the sequence of the protein and predicts the probability of the surface exposure for different regions.
EpiQuest-C is the program that analyses local immunological complexity of the protein sequence.
The higher is the complexity of the peptides, the higher is the probability of it being highly specific for this sequence only.
Predicting the relative strength of an epitope
The most critical difference between EpiQuest-B and all other B-epitope prediction software is that the Antigenicity Index that the program assigns to different peptide sequences directly correlates to the actual immunogenicity of the epitope (its ability to elicit an immune response.
Here we show data for 9 different peptide epitopes from the sequence of NS1 protein of Dengue virus 2. Twp different strains of mice (blue and red dots) were immunized either using peptide that includes all these epitopes, or recombinant NS1 or were infected with lice DV2. In all cases the antigenicity index assigned to every peptide correlated with the level of antibody titre directed against this particular sequence. Read more.




